MMTO Queue Catalog User Interface
PIs of MMT programs that will be observed in the MMIRS or Binospec queues manage their catalogs through a web interface. This page documents the catalog management. Most features of the web interface are easy to use; this page provides a reference, and documents some special cases.
Contents
- Accessing your catalog
- Catalog view
- Designing a multi-slit mask
- Creating or modifying a target
- Writing helpful observation notes
Accessing your catalog
Each program has an email contact address, and the contact person will get an email titled “MMT Observatory Queue Observing Form” from the MMT Scheduling System about 1-2 months before the observing run, with a link to log in to the catalog. You must receive and read these emails to be aware of submission deadlines and to be able to log into your catalog. If you can’t find the emails, try searching for and whitelisting “MMT Scheduler”, “MMT Scheduling System”, and “scheduler@mmto.org”.
Logging in takes you to a page that shows the deadlines for slitmask submission and catalog submission before the run, typically about 2 weeks and 1 week before the run starts for Binospec, but longer for MMIRS. You must check a box acknowledging the instructions and submit this form to access the catalog editor.
The mask and catalog deadlines are important to get your masks made and targets verified before the run starts. If your mask designs are not in on time, the masks cannot be machined and shipped to the MMT in time for the run (and MMIRS masks must be loaded into the mask dewar before it is cooled down). Even for imaging and longslit targets, your targets must be added to the catalog by the target deadline before the run so that the instrument scientist and queue observers can check for problems and run queue scheduling software. The catalog will lock before the observing run, and any further changes require the PI to request that MMT staff unlock the catalog. PIs of ToO or time-variable programs must contact the MMT instrument scientist to discuss their catalog, unlock requests, and the procedures for triggering a ToO.
If you want a collaborator to have access to your queue catalog to be able to edit targets and design masks, you can forward them the email with the catalog link, or just copy and send them the link URL containing a token string. Be careful with this link, since anyone with the link can edit that catalog.
Catalog view
The middle of the catalog page shows a table of your targets. To create new entries in the target table, use the new, copy, or upload buttons just above the table. In the upper right of the window is a list of masks that you have designed. There is a distinction between “target” and “mask.” A target defines an observation, so you can have two targets that use the same mask, for example, short and long exposure times, or observations in two different semesters.
When you click “copy” to copy an observation, there are some options to browse the current catalog, or your previous trimester’s catalogs, etc. Some of these previously had an option to copy “by reference” – to a pointer to the previous observation – this is grayed out now because it could cause problems. For most purposes, it is better to create a new copy (for example, if you have a partially completed observation from a previous trimester, and a new time allocation, you want to make a new copy).
Screenshot of the target table:
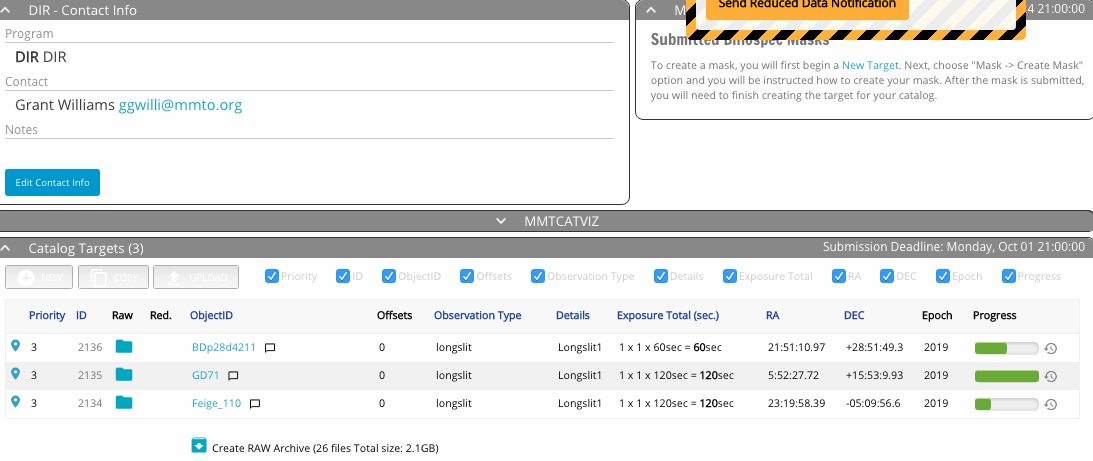
The queue catalog window showing the target table.
Below the catalog window is a summary of your requested time including estimated overheads. Overheads include wavefront sensing, acquisition time, readout, and calibrations. The queue observers will typically keep an accounting of the actual overhead time used during an observation, and charge that rather than the estimate.
You must assign a priority 1, 2, or 3 to each observation. These help us understand which observations are most important to you and prioritize data taking. Please assign priorities so that roughly 1/3rd of your time is in each category. If you have a small number of targets, splitting in thirds may be difficult – just try to assign priorities reasonably. We look at the overall completion percentage of a program to split time fairly among programs, so assigning priority 1 to all of your programs won’t get you more data, it just makes it harder to tell which targets are most important (so don’t do that). You don’t have to squeeze into the total times exactly, or choose unusual exposure times, to fit it precisely into the time allocation. In other words if you have 3.9 hours of priority 1 time, and you want to give 2 hours exposure to each of N targets, go ahead and make the two most important targets priority 1 with 2 hours each; you don’t have to make it 1.95 hours each.
Screenshot of the time allocation summary:
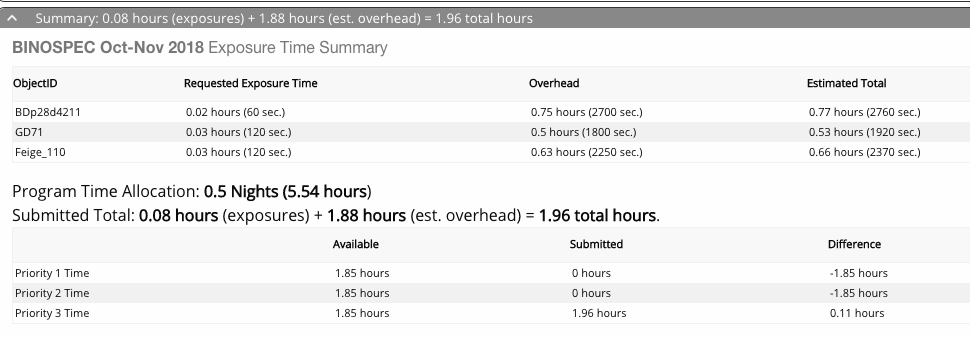
The lower part of the queue catalog window showing the time allocation summary.
The columns of the target table are mostly self-explanatory. A few useful icons include: clicking the location pin on the left will open a sky visualization of the target; when data is taken, folder and box icons appear in the Raw and Reduced columns for data download; and clicking the clock icon on the extreme right brings up the history of the observing block, including time of observation and observer’s notes.
Above the target table is a gray bar labeled “MMTCATVIZ.” Clicking this will expand a sky visualization of the location of the targets. To hide it, click the bar again.
Designing a slit mask
To design a new slit mask, click the “New Target” button/text either in the catalog panel or the mask list on upper right. This will open a new target wizard (series of dialog boxes). Click “mask” and then “create new mask.” This should open mask design software in the browser for Binospec; for MMIRS, you run the xfitmask mask design software as a standalone program. Descriptions of the mask designers are available separately for BinoMask and for MMIRS xfitmask. For BinoMask, the mask designer communicates the design directly to the queue database, and for MMIRS, you will run the design program and upload the files.
When done designing the mask, you return to the catalog page, click “New Target” again, and choose “Already Submitted or Existing Mask.” This will load the parameters of RA, Dec, PA from the mask design file, and then let you choose exposure time, etc to create an entry in the target table.
It’s a good idea to give your mask a memorable name, after your target, program, or PI, eg “ngc5894_v1_mask1” or “JSmith_field2”, etc. A unique ID number is appended to the mask name, but having masks with generic names e.g. “Mask3” makes it harder to keep track. Also, if you reuse masks in another semester you’ll be glad for informative names.
The queue catalog keeps track of masks that have been previously designed by a PI, so when you make a target with “Already Submitted or Existing Mask,” you should get a list that allows you to choose masks that you designed in previous semesters. If you need to find a mask that is not in the list, or if you need to use a mask designed by another person, contact the MMT instrument scientists.
Often, designing slitmasks takes some trial and error, and you may wind up designing a second version of your slitmasks. It helps to add a “_v2” suffix or similar to the names to keep track. If you iterate on mask designs, it’s not currently possible to delete the old mask design files, but please contact your MMT instrument scientist to request that the unnecessary masks not be machined.
Creating or modifying a target
To create a target, use the new, copy, or upload buttons above the table. Clicking on an existing target name in the table lets you edit it. For a new target, the catalog target wizard dialog boxes will appear, and prompt to choose one of the three types of targets: imaging, longslit, or slitmask. For a mask, you then choose from your designed masks, and the RA, Dec, and PA are read from the mask file. For imaging and longslit targets, you fill in the RA, Dec, and PA.
You will fill in typical catalog details: name, RA, Dec, and so on. Use an informative name that is likely to be distinct from other targets in the queue: eg “Virgo_dwarf_1” is better than “Virgo1”. If you copy a target, the software will generate a unique name by appending a long number. We strongly recommend editing your target names to some relatively short but unique set of names – long IDs that differ by only a few digits are harder for observers to read at 4 AM.
The catalog input boxes will show help text if input is not in the correct format. Some notes on input formats:
- RA and Dec must be in hh:mm:ss.s and dd:mm:ss.s.
- Priority should be 1, 2, or 3.
- Proper motions are not in mas/yr, but must be in PM_RA = seconds of time/century, and PM_Dec = seconds of arc/century. These can be converted by:
pm_RA (sec of time/century) = pm_RA(mas/yr) / [10 * 15 * cos(Dec)] pm_Dec (sec of arc/century) = pm_Dec(mas/yr) / 10
- PA is in degrees of East of North, as usual. If you want parallactic angle, write that in the notes. However, the f/5 optical instruments (Binospec and Hectospec) are behind an atmospheric dispersion compensator (ADC), so the parallactic angle is less relevant and you don’t need to request it, although we will often try to observe Binospec longslits near parallactic anyway.
- Magnitude is for the queue observer’s information, so they know e.g. whether the object is too faint to do in less-good conditions, and to determine if the data quality is good (looking at the images to see the flux expected for a given mag). Please give a magnitude for the most important science target, eg if your mask has a mag=20 primary target and fillers at mag 18-21, you should enter mag 20.
- Epoch is the time at which the RA/Dec coordinates are valid, e.g. 2015.5 for Gaia DR2. This matters for high proper-motion objects, including some standard stars. We have had an issue with the MMT mount code using this number for both epoch and equinox (epoch = time of observation; equinox = reference frame, which is normally J2000). As of 8/2022, precession in the mount code has been disabled, but there is still a bug in handling non-J2000 equinoxes. So, in practice entering a non-2000 epoch such as 2015.5 for Gaia DR2 still leads to problems. We recommend using standard stars with small proper motions whenever possible; if not, apply PMs to advance RA/Dec to the current epoch, and enter 0 for the PMs and 2000 for the epoch/equinox.
- For information about filters, gratings, wavelengths, etc, see the instrument webpages. If you have questions, contact your MMT instrument scientist.
The next dialog box sets the exposure time. The total exposure on your target will be
total = (exposure length) x (number of exposures) x (number of visits)
For example, if you want six hours on a single Binospec mask, you could enter 1200 sec exposures, 18 exposures, and 1 visit. For Binospec, 1200 sec is the longest recommended spectroscopic exposure time due to cosmic rays. Spectroscopic exposures should be less than 1200 sec, and imaging exposures less than 120 sec (even less in crowded fields, due to saturation and persistence). The queue software will break a long sequence up into 2-hour observing blocks for ease of scheduling. If you absolutely need contiguous observations, e.g. for a time sensitive observation such as a transit, put that and the time of the event in your target notes.
The overhead is estimated once per observing block. This means that if you have an exposure longer than 2 hours, you’ll get charged more than one overhead. For example, a 6 hour slitmask integration would incur 1.5 hours of overhead (3 OBs x 0.5 hours per OB). Thus, for example, putting in a lot of targets each with 2.5 hours integration will incur more overhead than is optimal – you could give the brighter ones 2 hours and the fainter ones 3 hours, reduce overhead and the number of OBs, and make the observations easier to schedule.
The purpose of the “visit” is to indicate splitting your exposure, e.g. for monitoring programs. So if you wanted to observe a target on six nights for one hour each night, you would specify 6 visits and one visit per night. For multiple visits or any other special constraint on observations, please give instructions in the target notes (like “Please take 3 visits each spaced 3 or more days apart”). If you don’t care about monitoring, you can enter 1 visit, like: 1 visit x 12 exposures x 1200 sec.
Special observation instructions including time constraints, weather/seeing constraints, and requesting dithering, flux standards, or twilight flats, should also go in the target notes.
Clicking on a target name in the target table will bring up a dialog box to edit the target parameters. This box also allows you to delete a target, if needed.
The target creation and edit windows allow you to specify offset stars and to upload a finding chart image. MMIRS can use offset stars; see the MMIRS pages for how to set the longslit at a given PA to include both science target and offset star. MMIRS also requires finding charts for longslit targets. Please note that Binospec doesn’t use offset stars or finding charts, but sets to your target directly; you must have coordinates on the Gaia DR2 reference system (SDSS is close enough) to accurately acquire the target. There is not a way for the Binospec observers to set onto your target with finding charts or offset stars, because there is no slit viewing camera, so your coordinates must be good.
Screenshot of the target editing dialog:
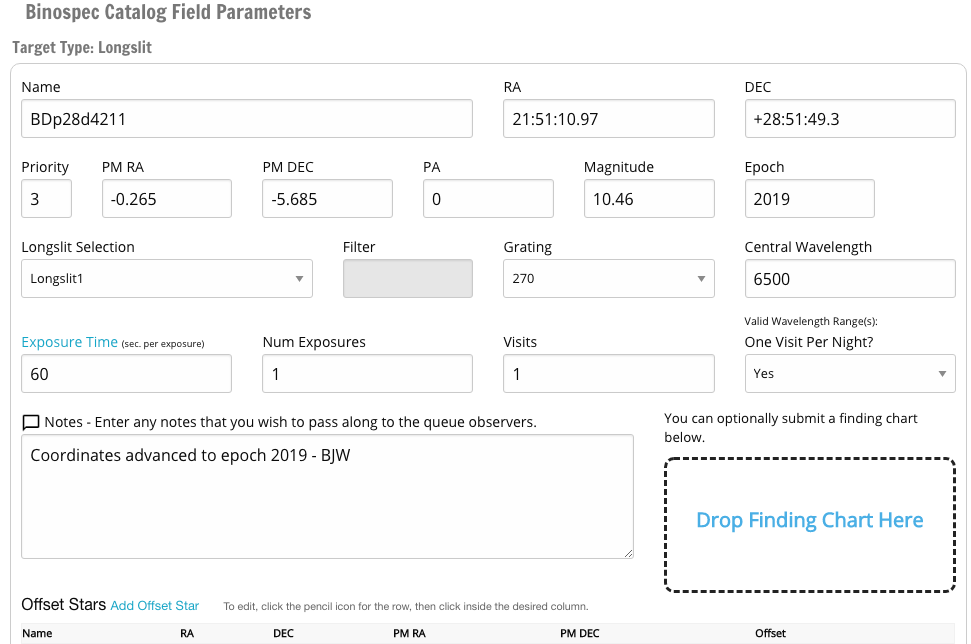
The dialog box that pops up to edit a target when you click on its name. The save/cancel/delete buttons and the finding chart can be reached by scrolling down.
Writing helpful observation notes
The observation notes/comment box should be used to convey information to the queue observers that the PI knows, but the observers will not be able to figure out from the target parameters alone. The notes should be concise. Please keep in mind that the observers are reading these notes in the middle of the night, and if you write a lot of text, the important parts may get overlooked.
Information that is helpful in the target notes includes for example: target priorities, amount of flexibility in observing parameters, unusual target properties, and time-series requests. It is not so helpful to attempt to micro-manage the observing procedure or to request overly stringent observing conditions. Basically, tell the staff what you want to accomplish, more than how to do it.
Some examples of helpful observing notes:
- “This is our highest priority target.”
- “Slit PA is along major axis of galaxy but can be varied by +/-5 deg if needed to find guide stars.”
- “These 6 visits are to sample a radial velocity curve of a binary with a period of 23 hours, please make one visit per night and stagger the time slightly later each night to sample the orbit.”
- “Please take a spectrum of the standard star in our catalog [such as Feige34_270_6500] at some point during the run for relative flux calibration.” Or “Please take a spectrum of the standard star in our catalog on the same night,” if you have strong needs for same-night calibration and want to spend the observing time on it.
- “Target is extremely red and will only show flux at long wavelengths.”
- “Spectrum of a large low surface brightness galaxy, ok to do in mediocre seeing.”
Examples of observing notes that are less helpful or tend to clutter up the notes:
- Extremely detailed offset or dither requests – we typically offset targets for longslit or imaging to be away from chip/amplifier boundaries, as appropriate for the instrument.
- Requesting Binospec or Hectospec observations at the parallactic angle – these instruments are behind an atmospheric dispersion compensator, so the parallactic angle matters less. For Binospec longslits, we will often try to observe near parallactic anyway. For Binospec slitmasks, longslits with a particular PA, or Hectospec, the sky PA is constrained and we can’t observe at parallactic.
- Blind offset stars for Binospec, which can’t use them (no slit viewing camera), but can set directly onto your target RA/Dec accurately.
- Blind offset stars with ra/dec offsets for MMIRS – the ra/dec offsets don’t work well in practice. You can specify a PA that puts the slit through both your target and a brighter offset star, provide a finding chart, and the observers can align the slit onto the star.
- Airmass or seeing requirements that make an observation hard to schedule. If you enter the correct target magnitude, we will observe in appropriate conditions, eg. we won’t observe mag=24 targets in 2″ seeing or clouds. If you request better than median conditions (eg better than 1″ seeing in the optical), the observers are reluctant to start your target if the seeing is at all variable, so you wind up getting less data.
- Constrictive scheduling requirements that are difficult or unfeasible, such as “Please observe the velocity curve over a single 12 hour period” (not possible in a single night); or “Please observe on 12 consecutive nights” (you have to add some flexibility since we are unlikely to get 12 clear nights in a row, not all instrument runs are 12 nights long, etc).
If you have questions about how to specify your observation, please always feel free to ask your instrument scientist – it is better to ask than to write an observation specification that is hard to schedule, and then wonder why you aren’t getting any data.
For questions about entering your targets and designing your observations, please contact your MMT instrument scientist: for Binospec and MMIRS, Benjamin Weiner. The Hectospec queue is managed independently of this interface by Nelson Caldwell at CfA. The MMT queue catalog management web pages were implemented by Dallan Porter.
Benjamin Weiner, bjw@mmto.org
